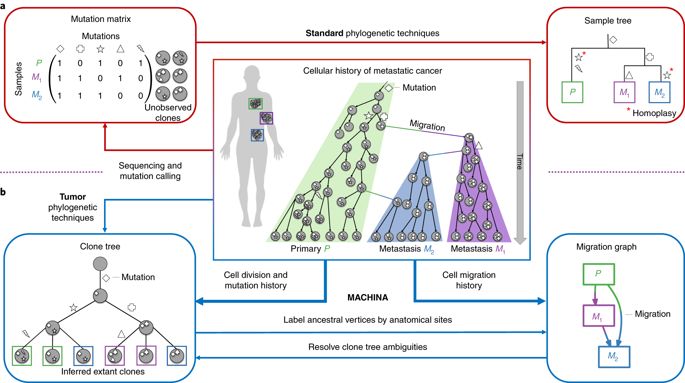
 Phylogenetic analysis of metastatic tumors. Source: Princeton/Nature Genetics
Phylogenetic analysis of metastatic tumors. Source: Princeton/Nature Genetics
Princeton researchers have developed a new computational method that can track the spread of cancer cells through a patient’s body. Cell migration is often the symptom that leads to metastatic disease, which is the cause of 90 percent of solid tumor deaths. Doctors and researchers need to understand metastatic disease in order to develop treatments that stop cancers from spreading in the body.
The new algorithm tracks cancer metastasis by integrating DNA sequence data with information on where cells are located in the body. The algorithm is named MACHINA, for metastatic and clonal history integrative analysis.
"Our algorithm enables researchers to infer the past process of metastasis from DNA sequence data obtained at the present time," said Ben Raphael, a professor of computer science at Princeton and the senior author of the new research. "The data sets we get these days are very complex, but complex data sets don't always require complex explanations.”
The algorithm provides researchers with the clearest picture of cancer cell migration histories yet. The algorithm develops this image by tracking the DNA sequences.
MACHINA traces the cells’ mutations and movements. The algorithm found that, in some patients, metastasis could be caused by fewer cellular migrations than previously thought.
For example, in one breast cancer patient, a past study said that metastatic disease in this patient was started by 14 separate migrations. The new algorithm and study have found that one secondary tumor in the patient’s lung jumpstarted the metastasis in only five migrations. MACHINA has also been used to analyze patterns in patients who had melanoma, ovarian and prostate cancers.
The algorithm has tools that help its accuracy. A model for the co-migration of genetically different cells that is based on experimental evidence was developed for the algorithm. This evidence claims that cells can travel in clusters to new areas of a patient’s body. The model accounts for DNA uncertainty data for sequencing mixtures of genetically distinct tumor cells and healthy cells.
The algorithm helps doctors and researchers overcome previous challenges due to difficult data.
"I predict this new method will be of widespread use to the genomics community and will shed new light on the most deadly phase of cancer evolution," Andrea Sottoriva, the Chris Rokos Fellow in Evolution and Cancer at The Institute of Cancer Research, London, said.
The algorithm is opening doors for many new methods to examine metastatic patterns in cancer patients.
Researchers say that the method could be further developed by using data from tumor DNA and tumor cells in the bloodstream.
The paper on this research was published in Nature Genetics.

